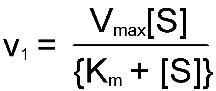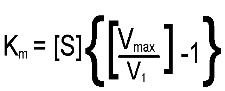Enzyme Kinetics
Introduction to Enzymes
Enzymes are biological catalysts responsible
for supporting almost all of the chemical reactions that maintain animal
homeostasis. Because of their role in maintaining life processes, the assay and
pharmacological regulation of enzymes have become key elements in clinical
diagnosis and therapeutics. The macromolecular components of almost all enzymes
are composed of protein, except for a class of RNA modifying catalysts known as
ribozymes. Ribozymes are molecules of ribonucleic
acid that catalyze reactions on the phosphodiester bond of other RNAs.
Enzymes are found in all tissues and fluids of the body. Intracellular
enzymes catalyze the reactions of metabolic pathways. Plasma membrane enzymes
regulate catalysis within cells in response to extracellular signals, and
enzymes of the circulatory system are responsible for regulating the clotting of
blood. Almost every significant life process is dependent on enzyme
activity.
back to the
top
Enzyme Classifications
Traditionally, enzymes were simply assigned names by the investigator who
discovered the enzyme. As knowledge expanded, systems of enzyme classification
became more comprehensive and complex. Currently enzymes are grouped into six
functional classes by the International Union of Biochemists (I.U.B.).
| No: |
Classification |
Biochemical
Properties |
| 1. |
Oxidoreductases |
Act on many chemical groupings to add or
remove hydrogen atoms. |
| 2. |
Transferases |
Transfer functional groups between donor and
acceptor molecules. Kinases are specialized transferases that regulate
metabolism by transferring phosphate from ATP to other
molecules. |
| 3. |
Hydrolases |
Add water across a bond, hydrolyzing
it. |
| 4. |
Lyases |
Add water, ammonia or carbon dioxide across
double bonds, or remove these elements to produce double bonds. |
| 5. |
Isomerases |
Carry out many kinds of isomerization: L to D
isomerizations, mutase reactions (shifts of chemical groups) and
others. |
| 6. |
Ligases |
Catalyze reactions in which two chemical
groups are joined (or ligated) with the use
of energy from ATP. |
These rules give each enzyme a unique number. The I.U.B. system also
specifies a textual name for each enzyme. The enzyme's name is comprised of the
names of the substrate(s), the product(s) and the enzyme's functional class.
Because many enzymes, such as alcohol dehydrogenase, are widely known in the
scientific community by their common names, the change to I.U.B.-approved
nomenclature has been slow. In everyday usage, most enzymes are still called by
their common name.
Enzymes are also classified on the basis of their composition. Enzymes
composed wholly of protein are known as simple
enzymes in contrast to complex enzymes,
which are composed of protein plus a relatively small organic molecule. Complex
enzymes are also known as holoenzymes. In this
terminology the protein component is known as the apoenzyme, while the non-protein component is known as the
coenzyme or prosthetic
group where prosthetic group describes a complex in
which the small organic molecule is bound to the apoenzyme by covalent bonds;
when the binding between the apoenzyme and non-protein components is
non-covalent, the small organic molecule is called a coenzyme. Many prosthetic groups and coenzymes are
water-soluble derivatives of vitamins. It
should be noted that the main clinical symptoms of dietary vitamin insufficiency
generally arise from the malfunction of enzymes, which lack sufficient cofactors
derived from vitamins to maintain homeostasis.
The non-protein component of an enzyme may be as simple as a metal ion or as
complex as a small non-protein organic molecule. Enzymes that require a metal in
their composition are known as metalloenzymes if
they bind and retain their metal atom(s) under all conditions, that is with very
high affinity. Those which have a lower affinity for metal ion, but still
require the metal ion for activity, are known as metal-activated enzymes.
back to the
top
Role of Coenzymes
The functional role of coenzymes is to act as transporters of chemical
groups from one reactant to another. The chemical groups carried can be as
simple as the hydride ion (H+ + 2e-) carried by NAD
or the mole of hydrogen carried by FAD; or they can be even more complex than
the amine (-NH2) carried by pyridoxal phosphate.
Since coenzymes are chemically changed as a consequence of enzyme action, it
is often useful to consider coenzymes to be a special class of substrates, or
second substrates, which are common to many
different holoenzymes. In all cases, the coenzymes donate the carried chemical
grouping to an acceptor molecule and are thus regenerated to their original
form. This regeneration of coenzyme and holoenzyme fulfills the definition of an
enzyme as a chemical catalyst, since (unlike the usual substrates, which are
used up during the course of a reaction) coenzymes are generally regenerated.
back to the
top
Enzyme Relative to Substrate Type
Although enzymes are highly specific for the kind of reaction they catalyze,
the same is not always true of substrates they attack. For example, while
succinic dehydrogenase (SDH) always catalyzes an
oxidation-reduction reaction and its substrate is invariably succinic acid,
alcohol dehydrogenase (ADH) always catalyzes oxidation-reduction
reactions but attacks a number of different alcohols, ranging from methanol to
butanol. Generally, enzymes having broad substrate specificity are most active
against one particular substrate. In the case of ADH, ethanol is the preferred
substrate.
Enzymes also are generally specific for a particular steric configuration
(optical isomer) of a substrate. Enzymes that attack D sugars will not attack
the corresponding L isomer. Enzymes that act on L amino acids will not employ
the corresponding D optical isomer as a substrate. The enzymes known as
racemases provide a striking exception to these generalities; in
fact, the role of racemases is to convert D isomers to L isomers and vice versa.
Thus racemases attack both D and L forms of their substrate.
As enzymes have a more or less broad range of substrate specificity, it
follows that a given substrate may be acted on by a number of different enzymes,
each of which uses the same substrate(s) and produces the same product(s). The
individual members of a set of enzymes sharing such characteristics are known as
isozymes. These are the products of genes that
vary only slightly; often, various isozymes of a group are expressed in
different tissues of the body. The best studied set of isozymes is the
lactate dehydrogenase (LDH) system. LDH is a tetrameric enzyme
composed of all possible arrangements of two different protein subunits; the
subunits are known as H (for heart) and M (for skeletal muscle). These subunits
combine in various combinations leading to 5 distinct isozymes. The all H
isozyme is characteristic of that from heart tissue, and the all M isozyme is
typically found in skeletal muscle and liver. These isozymes all catalyze the
same chemical reaction, but they exhibit differing degrees of efficiency. The
detection of specific LDH isozymes in the blood is highly diagnostic of tissue
damage such as occurs during cardiac infarct.
back to the
top
Enzyme-Substrate Interactions
The favored model of enzyme substrate interaction is known as the induced fit model. This model proposes that the initial
interaction between enzyme and substrate is relatively weak, but that these weak
interactions rapidly induce conformational changes in the enzyme that strengthen
binding and bring catalytic sites close to substrate bonds to be altered. After
binding takes place, one or more mechanisms of catalysis generates transition-
state complexes and reaction products. The possible mechanisms of catalysis are
four in number:
1. Catalysis by Bond Strain: In this form of catalysis, the induced
structural rearrangements that take place with the binding of substrate and
enzyme ultimately produce strained substrate bonds, which more easily attain the
transition state. The new conformation often forces substrate atoms and bulky
catalytic groups, such as aspartate and glutamate, into conformations that
strain existing substrate bonds.
2. Catalysis by Proximity and Orientation: Enzyme-substrate
interactions orient reactive groups and bring them into proximity with one
another. In addition to inducing strain, groups such as aspartate are frequently
chemically reactive as well, and their proximity and orientation toward the
substrate thus favors their participation in catalysis.
3. Catalysis Involving Proton Donors (Acids) and Acceptors (Bases):
Other mechanisms also contribute significantly to the completion of catalytic
events initiated by a strain mechanism, for example, the use of glutamate as a
general acid catalyst (proton donor).
4. Covalent Catalysis: In catalysis that takes place by covalent
mechanisms, the substrate is oriented to active sites on the enzymes in such a
way that a covalent intermediate forms between the enzyme or coenzyme and the
substrate. One of the best-known examples of this mechanism is that involving
proteolysis by serine proteases, which include
both digestive enzymes (trypsin, chymotrypsin, and
elastase) and several enzymes of the blood
clotting cascade. These proteases contain an active site serine whose R
group hydroxyl forms a covalent bond with a carbonyl carbon of a peptide bond,
thereby causing hydrolysis of the peptide bond.
back to the
top
Chemical Reactions and Rates
According to the conventions of biochemistry, the rate of a chemical
reaction is described by the number of molecules of reactant(s) that are
converted into product(s) in a specified time period. Reaction rate is always
dependent on the concentration of the chemicals involved in the process and on
rate constants that are characteristic of the reaction. For example, the
reaction in which A is converted to B is written as follows:
A ------> B
The rate of this reaction is expressed algebraically as either a decrease in
the concentration of reactant A:
-[A] = k[B]
or an increase in the concentration of product B:
[B] = k[A]
In the second equation (of the 3 above) the negative sign signifies a
decrease in concentration of A as the reaction progresses, brackets define
concentration in molarity and the k is known as a rate constant. Rate
constants are simply proportionality constants that provide a quantitative
connection between chemical concentrations and reaction rates. Each chemical
reaction has characteristic values for its rate constants; these in turn
directly relate to the equilibrium constant for
that reaction. Thus, reaction can be rewritten as an equilibrium expression in
order to show the relationship between reaction rates, rate constants and the
equilibrium constant for this simple case. The rate constant for the forward
reaction is defined as k+1 and the reverse as k-1.
At equilibrium the rate (v) of the forward reaction (A -----> B) is--- by
definition--- equal to that of the reverse or back reaction (B -----> A), a
relationship which is algebraically symbolized as:
vforward = vreverse
where, for the forward reaction:
vforward = k+1[A]
and for the reverse reaction:
vreverse = k-1[B]
In the above equations, k+1 and k-1 represent rate
constants for the forward and reverse reactions, respectively. The negative
subscript refers only to a reverse reaction, not to an actual negative value for
the constant. To put the relationships of the two equations into words, we state
that the rate of the forward reaction [vforward] is equal to the
product of the forward rate constant k+1 and the molar concentration
of A. The rate of the reverse reaction is equal to the product of the reverse
rate constant k-1 and the molar concentration of B.
At equilibrium, the rate of the forward reaction is equal to the rate of the
reverse reaction leading to the equilibrium
constant of the reaction and is expressed by:
[B]/[A] = k+1/k-1 = Keq
This equation demonstrates that the equilibrium constant for a chemical
reaction is not only equal to the equilibrium ratio of product and reactant
concentrations, but is also equal to the ratio of the characteristic rate
constants of the reaction.
back to the
top
Chemical Reaction Order
Reaction order refers to the number of molecules involved in forming a
reaction complex that is competent to proceed to product(s). Empirically, order
is easily determined by summing the exponents of each concentration term in the
rate equation for a reaction. A reaction characterized by the conversion of one
molecule of A to one molecule of B with no influence from any other reactant or
solvent is a first-order reaction. The exponent on
the substrate concentration in the rate equation for this type of reaction is 1.
A reaction with two substrates forming two products would a second-order reaction. However, the reactants in second-
and higher- order reactions need not be different chemical species. An example
of a second order reaction is the formation of ATP through the condensation of
ADP with orthophosphate:
ADP + H2PO4 <----> ATP +
H2O
For this reaction the forward reaction rate would be written as:
vforward =
k1[ADP][H2PO4]
back to the
top
Enzymes as Biological Catalysts
In cells and organisms most reactions are catalyzed by enzymes, which are
regenerated during the course of a reaction. These biological
catalysts are physiologically important because they speed up the
rates of reactions that would otherwise be too slow to support life. Enzymes
increase reaction rates--- sometimes by as much as one millionfold, but more
typically by about one thousand fold. Catalysts speed up the forward and reverse
reactions proportionately so that, although the magnitude of the rate constants
of the forward and reverse reactions is are increased, the ratio of the rate
constants remains the same in the presence or absence of enzyme. Since the
equilibrium constant is equal to a ratio of rate constants, it is apparent that
enzymes and other catalysts have no effect on the equilibrium constant of the
reactions they catalyze.
Enzymes increase reaction rates by decreasing the amount of energy required
to form a complex of reactants that is competent to produce reaction products.
This complex is known as the activated state or transition
state complex for the reaction. Enzymes and other catalysts
accelerate reactions by lowering the energy of the transition state. The free
energy required to form an activated complex is much lower in the catalyzed
reaction. The amount of energy required to achieve the transition state is
lowered; consequently, at any instant a greater proportion of the molecules in
the population can achieve the transition state. The result is that the reaction
rate is increased.
back to the
top
Michaelis-Menton Kinetics
In typical enzyme-catalyzed reactions, reactant and product concentrations
are usually hundreds or thousands of times greater than the enzyme
concentration. Consequently, each enzyme molecule catalyzes the conversion to
product of many reactant molecules. In biochemical reactions, reactants are
commonly known as substrates. The catalytic event that converts substrate to
product involves the formation of a transition state, and it occurs most easily
at a specific binding site on the enzyme. This site, called the catalytic site of the enzyme, has been evolutionarily
structured to provide specific, high-affinity binding of substrate(s) and to
provide an environment that favors the catalytic events. The complex that forms
when substrate(s) and enzyme combine is called the enzyme
substrate (ES) complex. Reaction products arise when the ES complex
breaks down releasing free enzyme.
Between the binding of substrate to enzyme, and the reappearance of free
enzyme and product, a series of complex events must take place. At a minimum an
ES complex must be formed; this complex must pass to the transition state (ES*);
and the transition state complex must advance to an enzyme product complex (EP).
The latter is finally competent to dissociate to product and free enzyme. The
series of events can be shown thus:
E + S <---> ES <---> ES* <---> EP <---> E +
P
The kinetics of simple reactions like that above were first characterized by
biochemists Michaelis and Menten. The concepts underlying their analysis of
enzyme kinetics continue to provide the cornerstone for understanding metabolism
today, and for the development and clinical use of drugs aimed at selectively
altering rate constants and interfering with the progress of disease states. The
Michaelis-Menten equation:
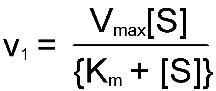
is a quantitative description of the relationship among the rate of an
enzyme- catalyzed reaction [v1], the concentration of substrate [S]
and two constants, Vmax and Km (which are set by the
particular equation). The symbols used in the Michaelis-Menton equation refer to
the reaction rate [v1], maximum reaction rate (Vmax),
substrate concentration [S] and the Michaelis-Menton constant (Km).
The Michaelis-Menten equation can be used to demonstrate that at the
substrate concentration that produces exactly half of the maximum reaction rate,
i.e.,1/2 Vmax, the substrate concentration is numerically equal to
Km. This fact provides a simple yet powerful bioanalytical tool that
has been used to characterize both normal and altered enzymes, such as those
that produce the symptoms of genetic diseases. Rearranging the Michaelis-Menton
equation leads to:
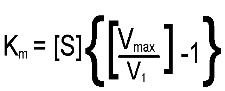 From this equation it should be apparent that when the substrate
concentration is half that required to support the maximum rate of reaction, the
observed rate, v1, will, be equal to Vmax divided by 2; in
other words, v1 = [Vmax/2]. At this substrate
concentration Vmax/v1 will be exactly equal to 2, with the
result that
From this equation it should be apparent that when the substrate
concentration is half that required to support the maximum rate of reaction, the
observed rate, v1, will, be equal to Vmax divided by 2; in
other words, v1 = [Vmax/2]. At this substrate
concentration Vmax/v1 will be exactly equal to 2, with the
result that
[S](1) = Km
The latter is an algebraic statement of the fact that, for enzymes of the
Michaelis-Menten type, when the observed reaction rate is half of the maximum
possible reaction rate, the substrate concentration is numerically equal to the
Michaelis-Menten constant. In this derivation, the units of Km are
those used to specify the concentration of S, usually Molarity.
The Michaelis-Menten equation has the same form as the equation for a
rectangular hyperbola; graphical analysis of reaction rate (v) versus substrate
concentration [S] produces a hyperbolic rate plot.
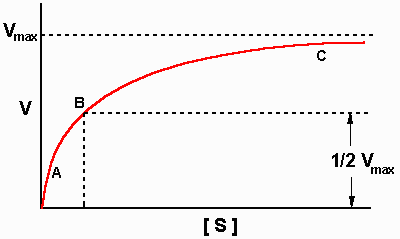 |
| Plot of substrate concentration
versus reaction velocity |
The key features of the plot are marked by points A, B and C. At high
substrate concentrations the rate represented by point C the rate of the
reaction is almost equal to Vmax, and the difference in rate at
nearby concentrations of substrate is almost negligible. If the Michaelis-Menten
plot is extrapolated to infinitely high substrate concentrations, the
extrapolated rate is equal to Vmax. When the reaction rate becomes
independent of substrate concentration, or nearly so, the rate is said to be
zero order. (Note that the reaction is zero order only with respect to this
substrate. If the reaction has two substrates, it may or may not be zero order
with respect to the second substrate). The very small differences in reaction
velocity at substrate concentrations around point C (near Vmax)
reflect the fact that at these concentrations almost all of the enzyme molecules
are bound to substrate and the rate is virtually independent of substrate, hence
zero order. At lower substrate concentrations, such as at points A and B, the
lower reaction velocities indicate that at any moment only a portion of the
enzyme molecules are bound to the substrate. In fact, at the substrate
concentration denoted by point B, exactly half the enzyme molecules are in an ES
complex at any instant and the rate is exactly one half of Vmax. At
substrate concentrations near point A the rate appears to be directly
proportional to substrate concentration, and the reaction rate is said to be
first order.
back to the
top
Inhibition of Enzyme Catalyzed Reactions
To avoid dealing with curvilinear plots of enzyme catalyzed reactions,
biochemists Lineweaver and Burk introduced an analysis of enzyme kinetics based
on the following rearrangement of the Michaelis-Menten equation:
[1/v] = [Km (1)/ Vmax[S] +
(1)/Vmax]
Plots of 1/v versus 1/[S] yield straight lines having a slope of
Km/Vmax and an intercept on the ordinate at
1/Vmax.
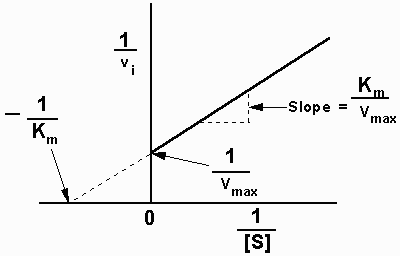 |
| A Lineweaver-Burk
Plot |
An alternative linear transformation of the Michaelis-Menten equation is the
Eadie-Hofstee transformation:
v/[S] = -v [1/Km] +
[Vmax/Km]
and when v/[S] is plotted on the y-axis versus v on the x-axis, the result is
a linear plot with a slope of -1/Km and the value
Vmax/Km as the intercept on the y-axis and Vmax
as the intercept on the x-axis.
Both the Lineweaver-Burk and Eadie-Hofstee transformation of the
Michaelis-Menton equation are useful in the analysis of enzyme inhibition. Since
most clinical drug therapy is based on inhibiting the activity of enzymes,
analysis of enzyme reactions using the tools described above has been
fundamental to the modern design of pharmaceuticals . Well- known examples of
such therapy include the use of methotrexate in cancer chemotherapy to
semi-selectively inhibit DNA synthesis of malignant cells, the use of aspirin to
inhibit the synthesis of prostaglandins which are at least partly responsible
for the aches and pains of arthritis, and the use of sulfa drugs to inhibit the
folic acid synthesis that is essential for the metabolism and growth of
disease-causing bacteria. In addition, many poisons--- such as cyanide, carbon
monoxide and polychlorinated biphenols (PCBs)--- produce their life- threatening
effects by means of enzyme inhibition.
Enzyme inhibitors fall into two broad classes: those causing irreversible
inactivation of enzymes and those whose inhibitory effects can be reversed.
Inhibitors of the first class usually cause an inactivating, covalent
modification of enzyme structure. Cyanide is a classic example of an
irreversible enzyme inhibitor: by covalently binding mitochondrial cytochrome
oxidase, it inhibits all the reactions associated with electron transport. The
kinetic effect of irreversible inhibitors is to decrease the concentration of
active enzyme, thus decreasing the maximum possible concentration of ES complex.
Since the limiting enzyme reaction rate is often k2[ES], it is clear
that under these circumstances the reduction of enzyme concentration will lead
to decreased reaction rates. Note that when enzymes in cells are only partially
inhibited by irreversible inhibitors, the remaining unmodified enzyme molecules
are not distinguishable from those in untreated cells; in particular, they have
the same turnover number and the same Km. Turnover
number, related to Vmax, is defined as the maximum number
of moles of substrate that can be converted to product per mole of catalytic
site per second. Irreversible inhibitors are usually considered to be poisons
and are generally unsuitable for therapeutic purposes.
Reversible inhibitors can be divided into two main categories--- competitive inhibitors and noncompetitive inhibitors---with a third category, uncompetitive inhibitors, rarely encountered.
| Inhibitor
Type |
Binding Site on
Enzyme |
Kinetic
effect |
| Competitive Inhibitor |
Specifically at the catalytic site, where it
competes with substrate for binding in a dynamic equilibrium- like
process. Inhibition is reversible by substrate. |
Vmax is unchanged; Km,
as defined by [S] required for 1/2 maximal activity, is
increased. |
| Noncompetitive Inhibitor |
Binds E or ES complex other than at the
catalytic site. Substrate binding unaltered, but ESI complex cannot form
products. Inhibition cannot be reversed by substrate. |
Km appears unaltered;
Vmax is decreased proportionately to inhibitor
concentration. |
| Uncompetitive Inhibitor |
Binds only to ES complexes at locations other
than the catalytic site. Substrate binding modifies enzyme structure,
making inhibitor- binding site available. Inhibition cannot be reversed by
substrate. |
Apparent Vmax decreased;
Km, as defined by [S] required for 1/2 maximal activity, is
decreased. |
The hallmark of all the reversible inhibitors is that when the inhibitor
concentration drops, enzyme activity is regenerated. Usually these inhibitors
bind to enzymes by non-covalent forces and the inhibitor maintains a reversible
equilibrium with the enzyme. The equilibrium constant for the dissociation of
enzyme inhibitor complexes is known as KI:
KI = [E][I]/[E--I--complex]
The importance of KI is that in all enzyme reactions where
substrate, inhibitor and enzyme interact, the normal Km and or
Vmax for substrate enzyme interaction appear to be altered. These
changes are a consequence of the influence of KI on the overall rate
equation for the reaction. The effects of KI are best observed in
Lineweaver-Burk plots.
Probably the best known reversible inhibitors are competitive inhibitors,
which always bind at the catalytic or active site of the enzyme. Most drugs that
alter enzyme activity are of this type. Competitive inhibitors are especially
attractive as clinical modulators of enzyme activity because they offer two
routes for the reversal of enzyme inhibition, while other reversible inhibitors
offer only one. First, as with all kinds of reversible inhibitors, a decreasing
concentration of the inhibitor reverses the equilibrium shown in reaction 6.8,
regenerating active free enzyme. Second, since substrate and competitive
inhibitors both bind at the same site, they compete with one another for binding
Raising the concentration of substrate (S), while holding the concentration
of inhibitor constant, provides the second route for reversal of competitive
inhibition. The greater the proportion of substrate, the greater the proportion
of enzyme present in competent ES complexes.
As noted earlier, high concentrations of substrate can displace virtually
all competitive inhibitor bound to active sites. Thus, it is apparent that
Vmax should be unchanged by competitive inhibitors. This
characteristic of competitive inhibitors is reflected in the identical
vertical-axis intercepts of Lineweaver-Burk plots, with and without inhibitor.
 |
| Lineweaver-Burk Plots of
Inhibited Enzymes |
Since attaining Vmax requires appreciably higher substrate
concentrations in the presence of competitive inhibitor, Km (the
substrate concentration at half maximal velocity) is also higher, as
demonstrated by the differing negative intercepts on the horizontal axis in
panel B.
Analogously, panel C illustrates that noncompetitive inhibitors appear to
have no effect on the intercept at the x-axis implying that noncompetitive
inhibitors have no effect on the Km of the enzymes they inhibit.
Since noncompetitive inhibitors do not interfere in the equilibration of enzyme,
substrate and ES complexes, the Km's of Michaelis-Menten type enzymes
are not expected to be affected by noncompetitive inhibitors, as demonstrated by
x-axis intercepts in panel C. However, because complexes that contain inhibitor
(ESI) are incapable of progressing to reaction products, the effect of a
noncompetitive inhibitor is to reduce the concentration of ES complexes that can
advance to product. Since Vmax = k2[Etotal],
and the concentration of competent Etotal is diminished by the amount
of ESI formed, noncompetitive inhibitors are expected to decrease
Vmax, as illustrated by the y-axis intercepts in panel C.
A corresponding analysis of uncompetitive inhibition leads to the
expectation that these inhibitors should change the apparent values of
Km as well as Vmax. Changing both constants leads to
double reciprocal plots, in which intercepts on the x and y axes are
proportionately changed; this leads to the production of parallel lines in
inhibited and uninhibited reactions.
back to the
top
Regulation of Enzyme Activity
While it is clear that enzymes are responsible for the catalysis of almost
all biochemical reactions, it is important to also recognize that rarely, if
ever, do enzymatic reactions proceed in isolation. The most common scenario is
that enzymes catalyze individual steps of multi-step metabolic pathways, as is
the case with glycolysis, gluconeogenesis
or the synthesis of
fatty acids. As a consequence of these lock- step sequences of reactions,
any given enzyme is dependent on the activity of preceding reaction steps for
its substrate.
In humans, substrate concentration is dependent on food supply and is not
usually a physiologically important mechanism for the routine regulation of
enzyme activity. Enzyme concentration, by contrast, is continually modulated in
response to physiological needs. Three principal mechanisms are known to
regulate the concentration of active enzyme in tissues:
- 1. Regulation of gene expression controls the quantity and rate of
enzyme synthesis.
- 2. Proteolytic enzyme activity determines the rate of enzyme
degradation.
- 3. Covalent modification of preexisting pools of inactive
proenzymes produces active enzymes.
Enzyme synthesis and proteolytic degradation are comparatively slow
mechanisms for regulating enzyme concentration, with response times of hours,
days or even weeks. Proenzyme activation is a more rapid method of increasing
enzyme activity but, as a regulatory mechanism, it has the disadvantage of not
being a reversible process. Proenzymes are generally synthesized in abundance,
stored in secretory granules and covalently activated upon release from their
storage sites. Examples of important proenzymes include pepsinogen, trypsinogen
and chymotrypsinogen, which give rise to the proteolytic digestive enzymes.
Likewise, many of the proteins involved in the cascade of chemical reactions
responsible for blood clotting are synthesized as proenzymes. Other important
proteins, such as peptide hormones and collagen, are also derived by covalent
modification of precursors.
Another mechanism of regulating enzyme activity is to sequester enzymes in
compartments where access to their substrates is limited. For example, the
proteolysis of cell proteins and glycolipids by enzymes responsible for their
degradation is controlled by sequestering these enzymes within the lysosome.
In contrast to regulatory mechanisms that alter enzyme concentration, there
is an important group of regulatory mechanisms that do not affect enzyme
concentration, are reversible and rapid in action, and actually carry out most
of the moment- to- moment physiological regulation of enzyme activity. These
mechanisms include allosteric regulation, regulation by reversible covalent
modification and regulation by control proteins such as calmodulin. Reversible
covalent modification is a major mechanism for the rapid and transient
regulation of enzyme activity. The best examples, again, come from studies on
the regulation of glycogen metabolism where phosphorylation of glycogen
synthase and glycogen phosphorylase kinase results in the
stimulation of glycogen degradation while glycogen synthesis is coordinately
inhibited. Numerous other enzymes of intermediary metabolism are affected by
phosphorylation, either positively or negatively. These covalent
phosphorylations can be reversed by a separate sub-subclass of enzymes known as
phosphatases. Recent research has indicated that the aberrant
phosphorylation of growth factor and hormone receptors, as well as of proteins
that regulate cell division, often leads to unregulated cell growth or cancer.
The usual sites for phosphate addition to proteins are the serine, threonine and
tyrosine R group hydroxyl residues.
back to the
top
Allosteric Enzymes
In addition to simple enzymes that interact only with substrates and
inhibitors, there is a class of enzymes that bind small, physiologically
important molecules and modulate activity in ways other than those described
above. These are known as allosteric enzymes; the
small regulatory molecules to which they bind are known as effectors. Allosteric effectors bring about catalytic
modification by binding to the enzyme at distinct allosteric sites, well removed
from the catalytic site, and causing conformational changes that are transmitted
through the bulk of the protein to the catalytically active site(s).
The hallmark of effectors is that when they bind to enzymes, they alter the
catalytic properties of an enzyme's active site. Those that increase catalytic
activity are known as positive effectors. Effectors that reduce or inhibit
catalytic activity are negative effectors.
Most allosteric enzymes are oligomeric (consisting of multiple subunits);
generally they are located at or near branch points in metabolic pathways, where
they are influential in directing substrates along one or another of the
available metabolic paths. The effectors that modulate the activity of these
allosteric enzymes are of two types. Those activating and inhibiting effectors
that bind at allosteric sites are called heterotropic
effectors. (Thus there exist both positive and negative heterotropic
effectors.) These effectors can assume a vast diversity of chemical forms,
ranging from simple inorganic molecules to complex nucleotides such as cyclic
adenosine monophosphate (cAMP). Their single defining feature is that they are
not identical to the substrate.
In many cases the substrate itself induces distant allosteric effects when
it binds to the catalytic site. Substrates acting as effectors are said to be
homotropic effectors. When the substrate is the
effector, it can act as such, either by binding to the substrate-binding site,
or to an allosteric effector site. When the substrate binds to the catalytic
site it transmits an activity-modulating effect to other subunits of the
molecule. Often used as the model of a homotropic effector is hemoglobin,
although it is not a branch-point enzyme and thus does not fit the definition on
all counts.
There are two ways that enzymatic activity can be altered by effectors: the
Vmax can be increased or decreased, or the Km can be
raised or lowered. Enzymes whose Km is altered by effectors are said
to be K-type enzymes and the effector a K-type
effector. If Vmax is altered, the enzyme and effector are said to be
V-type. Many allosteric enzymes respond to multiple effectors with V-type and
K-type behavior. Here again, hemoglobin is often used as a model to study
allosteric interactions, although it is not strictly an enzyme.
In the preceding discussion we assumed that allosteric sites and catalytic
sites were homogeneously present on every subunit of an allosteric enzyme. While
this is often the case, there is another class of allosteric enzymes that are
comprised of separate catalytic and regulatory subunits. The archetype of this
class of enzymes is cAMP-dependent protein kinase (PKA), whose
mechanism of activation is illustrated. The
enzyme is tetrameric, containing two catalytic subunits and two regulatory
subunits, and enzymatically inactive. When intracellular cAMP levels rise, one
molecule of cAMP binds to each regulatory subunit, causing the tetramer to
dissociate into one regulatory dimer and two catalytic monomers. In the
dissociated form, the catalytic subunits are fully active; they catalyze the
phosphorylation of a number of other enzymes, such as those involved in
regulating glycogen metabolism. The regulatory subunits have no catalytic
activity.
back to the
top
Back to Topics<<<<
This article has been modified by Dr. M. Javed Abbas.
If you have any comments please do not hesitate to sign my Guest Book.
20:38 21/12/2002
|